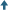PFAS present throughout the Yadkin-Pee Dee river food chain
Researchers have found per- and polyfluoroalkyl substances (PFAS) in every step of the Yadkin-Pee Dee River food chain, even though the river does not have a known industrial input of these compounds.
Gazette’s Introductory Note: This North Carolina State University research adds a new dimension to PFAS contamination of the environment. You can protect yourself from PFAS in drinking water with a home reverse osmosis unit, but it may be harder to avoid PFAS-contaminated foods. The ubiquitousness of PFAS especially brings into question the wisdom of the recent administration rejection of proposed EPA rules designed to limit PFAS content of imported goods.
Researchers from North Carolina State University have found per- and polyfluoroalkyl substances (PFAS) in every step of the Yadkin-Pee Dee River food chain, even though the river does not have a known industrial input of these compounds. The study examined the entire aquatic ecosystem for PFAS compounds and identified strong links between ecosystem groups that lead to biomagnification, the process that leads to greater concentrations of these substances in animals that sit higher on the food chain — including humans.
PFAS compounds were engineered to resist friction and heat, and are in many products that we use daily, from furniture to meat packaging. However, it is this “slippery” characteristic that makes them persist in ecosystems and poses a risk to our health.
“These compounds are engineered to be persistent on purpose; this is how they keep stains off your couch and eggs from sticking to your frying pan,” says Tom Kwak, unit leader of NC Cooperative Fish and Wildlife Research Unit, professor of applied ecology at NC State, and a co-author of the study. “We pay the price for these compounds when they enter the aquatic ecosystem.”
In a study measuring real-time PFAS contamination levels along the entire food chain of this major Atlantic river — from water and sediment to insects and fish — the researchers identified two PFAS hot spots along the Pee Dee and were able to establish strong links of PFAS transmission up the aquatic food chain.
The research team collected water, sediment, algae, plant, insect, fish, crayfish, and mollusk samples at five study sites along the length of the Yadkin-Pee Dee River, which begins in Blowing Rock, N.C., and runs 230 miles to empty into the Atlantic Ocean at Winyah Bay, South Carolina. They analyzed the samples for 14 different PFAS compounds.
While nearly every sample contained PFAS compounds, the site with the greatest PFAS concentrations was just downstream of the Rocky River input, which drains part of the watershed from Charlotte, N.C. and the surrounding area. The site with the second greatest PFAS concentrations was downstream in South Carolina, but there is no known or plausible input of PFAS for that region.
In aquatic food chains, biofilm — the soupy mixture of algae and bacteria that sticks to your boat — is the base resource for all life further up the chain. In this study, the largest concentrations of 10 of the 14 PFAS compounds measured were in biofilm samples. Unsurprisingly, aquatic insects, which primarily eat biofilm, had the greatest accumulation of PFAS compounds of all the living taxa the researchers sampled. This confirms a strong trophic link, or step in the food chain, showing how PFAS transfers from biofilm to insects, which are then eaten by freshwater fish.
When PFAS is in every step of the food chain, the compounds accumulate at each step. For example, a fish caught in an area with PFAS may have eaten hundreds of insects, each of which has consumed contaminated biofilm and other plants.
“We are part of the food chain and when we ingest these foods, we accumulate their PFAS loads, too,” says Greg Cope, William Neal Reynolds Distinguished Professor of Applied Ecology, coordinator of the NC State Agromedicine Institute, and corresponding author of the study. “This gives new meaning to the phrase, ‘You are what you eat.'”
Science Daily. (Report is slightly truncated.)